Plenary Speaker
 Richard Lenski
Richard Lenski
Hannah Distinguished Professor
Department of Microbiology & Molecular Genetics, Michigan State University
The dynamics of phenotypic and genomic evolution during a 50,000 generation experiment with E. coli
Twelve initially identical populations of E. coli have been propagated in a simple environment since 1988. Two goals of this long-term experiment have been to investigate the dynamics of evolutionary change and examine the reproducibility of outcomes. We have quantified the extent of adaptation by natural selection, documented increasing ecological specialization over time, observed the rise of mutator phenotypes, and even seen the origin of a new function that transcends the usual definition of E. coli as a species. We have pursued various genetic approaches to discover the mutations responsible for these changes, including several affecting global regulatory networks. We have also recently sequenced complete genomes to find all of the mutations present in temporal series of clones sampled from some populations and heterogeneous population samples to discover polymorphisms. These genomic data provide new insights into the dynamical coupling of phenotypic and genomic evolution, and into the role of complex mutations in the origin of new functions.
 Aimee Dunlap
Aimee Dunlap
Postdoctoral Fellow, Department of Ecology and Evolutionary Biology and the Center for Insect Science, University of Arizona
Experimental evolution of learning abilities: uncertainty and reliability
Why are some behaviors learned while others are innate? Why are some things learned more easily than others? Both questions are thought to be tied to patterns of environmental change, but experiments are lacking. I used statistical decision theory to model behaviors and fitness consequences, and experimental evolution studies with fruit flies where I manipulated patterns of environmental change across evolutionary time, to address both of these fundamental questions in learning. The first experiment tested the effects of the reliability of experience and the fixity of the best action upon the evolution of learning and non-learning in 30 generations. I found that indeed, the interaction of these two types of change determine when learning, and when non-learning evolve. The second study was a full factorial experiment manipulating the reliabilities of two modes of stimuli: olfactory and visual. After 40 generations, I found that as predicted, flies in environments where olfactory stimuli are reliable learned better about olfactory than color stimuli, with the same being true for color stimuli. These novel studies show the importance of reliability and change in evolution of learning and the usefulness of experimental evolution approaches to learning.
 Jenna Gallie
Jenna Gallie
Ph.D. Candidate in evolutionary genetics, New Zealand Institute for Advanced Study, Massey University.
Experimental evolution of bet hedging
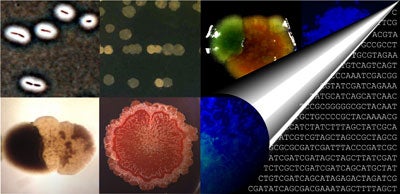 Bet hedging – stochastic switching between phenotypic states – is a canonical example of an evolutionary adaptation that facilitates persistence in the face of fluctuating environments. Although bet hedging is found in organisms ranging from bacteria to humans, direct evidence for an adaptive origin of this behaviour is lacking. Here I report the de novo evolution of bet hedging in experimental bacterial populations. Bacteria were subjected to an environment that continually favoured new phenotypic states. Initially, this regime drove the successive evolution of novel phenotypes by mutation and selection; however, in two (of 12) replicates this trend was broken by the evolution of bet-hedging genotypes that persisted because of rapid stochastic phenotype switching. Genome re-sequencing of one of these switching types revealed nine mutations that distinguished it from the ancestor. The final mutation proved both necessary and sufficient for rapid phenotype switching; nonetheless, the evolution of bet hedging was contingent upon earlier mutations that altered the relative fitness effect of the final mutation. I will account for the adaptive significance of each of the nine mutations and describe the effect of the final mutation that establishes a bi-stable (epigenetic) molecular switch. Together, these findings capture the adaptive evolution of bet hedging in the simplest of organisms, and suggest that risk-spreading strategies may have been among the earliest evolutionary solutions to life in fluctuating environments.
Bet hedging – stochastic switching between phenotypic states – is a canonical example of an evolutionary adaptation that facilitates persistence in the face of fluctuating environments. Although bet hedging is found in organisms ranging from bacteria to humans, direct evidence for an adaptive origin of this behaviour is lacking. Here I report the de novo evolution of bet hedging in experimental bacterial populations. Bacteria were subjected to an environment that continually favoured new phenotypic states. Initially, this regime drove the successive evolution of novel phenotypes by mutation and selection; however, in two (of 12) replicates this trend was broken by the evolution of bet-hedging genotypes that persisted because of rapid stochastic phenotype switching. Genome re-sequencing of one of these switching types revealed nine mutations that distinguished it from the ancestor. The final mutation proved both necessary and sufficient for rapid phenotype switching; nonetheless, the evolution of bet hedging was contingent upon earlier mutations that altered the relative fitness effect of the final mutation. I will account for the adaptive significance of each of the nine mutations and describe the effect of the final mutation that establishes a bi-stable (epigenetic) molecular switch. Together, these findings capture the adaptive evolution of bet hedging in the simplest of organisms, and suggest that risk-spreading strategies may have been among the earliest evolutionary solutions to life in fluctuating environments.
 William Harcombe
William Harcombe
Postdoctoral Researcher, Organismic and Evolutionary Biology Department, Harvard University
Microbial community dynamics: a systems biology and experimental evolutionary approach
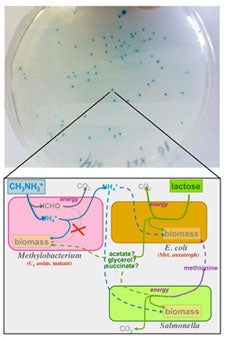 Most organisms exist as members of complex communities. Experimental evolution has provided important insight into how selection acts on genetic variation to drive adaptation in monocultures, however less is known about the dynamics of adaptation in a community context. How rugged are fitness landscapes when fitness is determined by species interactions? I investigate the evolution of model microbial communities in which species rely on one another for essential metabolites, i.e. cross-feed. Specifically, I study two and three-species consortia containing Escherichia coli, Salmonella typhimurium and Methylobacterium extorquens. Evolution dramatically altered the productivity of cross-feeding communities depending on the scale of species interactions. Adaptation in an environment with local interactions led to the evolution of novel interspecies cooperation; Salmonella evolved to secrete a costly amino acid needed by E. coli. Community members are being swapped between replicate evolved communities to determine the degree of co-evolution between partners. Additionally, genomic, transcriptomic, and metabolomic analyses are providing insight into the systems level mechanisms behind changes in community phenotype. Finally, genome level metabolic networks are being used to computationally predict the possible and optimal species interactions based on first principles of metabolic stoichiometry. This work is providing insight into how evolution shapes the emergent properties of consortia through altering the underlying species interactions, cellular metabolism, and genomic content of populations.
Most organisms exist as members of complex communities. Experimental evolution has provided important insight into how selection acts on genetic variation to drive adaptation in monocultures, however less is known about the dynamics of adaptation in a community context. How rugged are fitness landscapes when fitness is determined by species interactions? I investigate the evolution of model microbial communities in which species rely on one another for essential metabolites, i.e. cross-feed. Specifically, I study two and three-species consortia containing Escherichia coli, Salmonella typhimurium and Methylobacterium extorquens. Evolution dramatically altered the productivity of cross-feeding communities depending on the scale of species interactions. Adaptation in an environment with local interactions led to the evolution of novel interspecies cooperation; Salmonella evolved to secrete a costly amino acid needed by E. coli. Community members are being swapped between replicate evolved communities to determine the degree of co-evolution between partners. Additionally, genomic, transcriptomic, and metabolomic analyses are providing insight into the systems level mechanisms behind changes in community phenotype. Finally, genome level metabolic networks are being used to computationally predict the possible and optimal species interactions based on first principles of metabolic stoichiometry. This work is providing insight into how evolution shapes the emergent properties of consortia through altering the underlying species interactions, cellular metabolism, and genomic content of populations.
 Ellie Harrison
Ellie Harrison
Sex and selfishness: virulence evolution in a genetic parasite
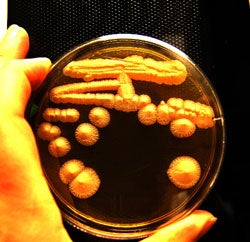 Genetic elements that are transmitted in a non-Mendelian manner can be subject to within-individual selection as well as the more conventional between-individual selection. The magnitude of this within-individual selection depends on the breeding system, being larger in outcrossed species and smaller in inbred or asexual species. We have studied the effect of varying the magnitude of the within-individual component of selection by analyzing the evolution of the 2 micron plasmid in experimental sexual and asexual populations of Saccharomyces cerevisiae. The 2 micron is a biparentally transmitted nuclear plasmid present in ~40 copies per haploid cell and appears to be entirely parasitic. In an asexual population plasmid fitness is entirely linked to that of its host, and selection to minimize harm is strong. However, in sexually reproducing populations the parasite is able to spread, and host imposed selection is reduced. As predicted, we find that the plasmid evolves higher copy number in the sexual populations. To our knowledge, this is the first experimental demonstration of parasite virulence evolving in response to the host sexual system.
Genetic elements that are transmitted in a non-Mendelian manner can be subject to within-individual selection as well as the more conventional between-individual selection. The magnitude of this within-individual selection depends on the breeding system, being larger in outcrossed species and smaller in inbred or asexual species. We have studied the effect of varying the magnitude of the within-individual component of selection by analyzing the evolution of the 2 micron plasmid in experimental sexual and asexual populations of Saccharomyces cerevisiae. The 2 micron is a biparentally transmitted nuclear plasmid present in ~40 copies per haploid cell and appears to be entirely parasitic. In an asexual population plasmid fitness is entirely linked to that of its host, and selection to minimize harm is strong. However, in sexually reproducing populations the parasite is able to spread, and host imposed selection is reduced. As predicted, we find that the plasmid evolves higher copy number in the sexual populations. To our knowledge, this is the first experimental demonstration of parasite virulence evolving in response to the host sexual system.
 Sara Mitri
Sara Mitri
Experimental evolution in communicating robots
 All social organisms use communicative signals to coordinate their behaviors with members of their own and other species. Despite this key role in social organization, the conditions conducive to the evolution of communication and the paths by which reliable systems of communication become established remain largely unknown. This is in part due to the difficulty of conducting experimental evolution on social species. In this talk, I will show how we circumvent this problem by using a system of experimental evolution with groups of foraging robots that could emit and perceive light to communicate.
All social organisms use communicative signals to coordinate their behaviors with members of their own and other species. Despite this key role in social organization, the conditions conducive to the evolution of communication and the paths by which reliable systems of communication become established remain largely unknown. This is in part due to the difficulty of conducting experimental evolution on social species. In this talk, I will show how we circumvent this problem by using a system of experimental evolution with groups of foraging robots that could emit and perceive light to communicate.
Our experiments revealed that the reliability of the resulting communication system depends on the level of relatedness between robots in a group. Robots that were highly related evolved reliable signals. In contrast, when relatedness between robots in a group was low, robots were selected to suppress the inadvertent cues produced while foraging. However, because of the effect of mutations, these cues were never completely suppressed and some variability in signaling was maintained. Because similar co-evolutionary processes should be common in natural systems, our findings predict that relatedness will play an important role in the evolution of natural systems of communication.
 Levi Morran
Levi Morran
Why sex with a partner is better: mutation load and rapid adaptation favor outcrossing over self-fertilization
 Outcrossing is the most common form of sexual reproduction among animals and plants. Despite this prevalence, the selective pressures maintaining the widespread use of outcrossing are not fully understood. Evolutionary theory predicts that outcrossing populations should be less susceptible to the fixation of deleterious mutations and may more rapidly adapt to changing ecological conditions than self-fertilizing populations. We empirically tested these predictions using experimental evolution, by exposing obligate selfing, mixed mating, and obligate outcrossing populations of C. elegans to elevated mutation rates, rearing them in a selective novel environment, and exposing them to a virulent pathogen. After fifty generations of mutation and selection, outcrossing populations fixed significantly fewer deleterious mutations than selfing populations and exhibited significantly greater rates of adaptation under natural mutation rates. Additionally, outcrossing populations rapidly adapted to the pathogen, whereas selfing populations failed to adapt. This work is the first empirical test of theory regarding the evolution and maintenance of outcrossing and demonstrates that, as predicted, outcrossing is conditionally favored over self-fertilization. However, the conditional value of outcrossing may be quite significant as most organisms are presumably subject to deleterious mutations and/or environmental conditions favoring rapid adaptation. Both of these factors likely explain the prevalence of outcrossing as a sexual mating strategy.
Outcrossing is the most common form of sexual reproduction among animals and plants. Despite this prevalence, the selective pressures maintaining the widespread use of outcrossing are not fully understood. Evolutionary theory predicts that outcrossing populations should be less susceptible to the fixation of deleterious mutations and may more rapidly adapt to changing ecological conditions than self-fertilizing populations. We empirically tested these predictions using experimental evolution, by exposing obligate selfing, mixed mating, and obligate outcrossing populations of C. elegans to elevated mutation rates, rearing them in a selective novel environment, and exposing them to a virulent pathogen. After fifty generations of mutation and selection, outcrossing populations fixed significantly fewer deleterious mutations than selfing populations and exhibited significantly greater rates of adaptation under natural mutation rates. Additionally, outcrossing populations rapidly adapted to the pathogen, whereas selfing populations failed to adapt. This work is the first empirical test of theory regarding the evolution and maintenance of outcrossing and demonstrates that, as predicted, outcrossing is conditionally favored over self-fertilization. However, the conditional value of outcrossing may be quite significant as most organisms are presumably subject to deleterious mutations and/or environmental conditions favoring rapid adaptation. Both of these factors likely explain the prevalence of outcrossing as a sexual mating strategy.
 Bjørn Østman
Bjørn Østman
Impact of epistasis and pleiotropy on evolutionary adaptation
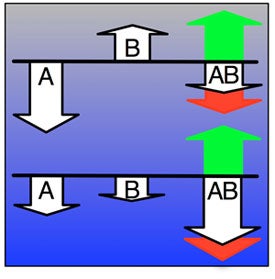 We investigate the impact of epistasis and pleiotropy on adaptive evolution in an NK model, by evolving a population of asexual haploid organisms with circular binary genomes of length N, where each locus interacts with K neighbors. We use a quantitative measure of the magnitude of epistatic interactions (size of epistasis) and find that it is a monotonically increasing function of K. At high mutation rates more epistatic pairs are observed on the line of descent than expected, whereas at low mutation rates some epistatic interactions are selected against. Higher fitness is attained at an intermediate amount of ruggedness because both the height of the global peak and the mean selection coefficient per beneficial mutation increase with K. However, if K is too high, the longer waiting time between beneficial mutations counteracts those benefits. Increasing K transforms the fitness landscape from non-epistatic and smooth to epistatic and rugged, and increases the adaptive potential of the population. When the environment changes so that the population finds itself at a suboptimal location of an epistatic landscape, pleiotropy enables the population to increase fitness drastically with only a few mutational steps. Subsequently, crossing valleys between peaks in the fitness landscape requires deleterious mutations, which then become essential for adaptation by interacting with later mutations. Therefore, both deleterious and beneficial mutations contribute to adaptation, underscoring the benefit of initially deleterious mutations and epistasis in adaptive evolution. Increasing the number of interactions between loci on the one hand creates the opportunity for groups of loci to work synergistically as modules towards increased fitness. On the other hand, if the number of interactions is too high, the constraints they engender may be too severe for the adaptive process to take advantage of the potential.
We investigate the impact of epistasis and pleiotropy on adaptive evolution in an NK model, by evolving a population of asexual haploid organisms with circular binary genomes of length N, where each locus interacts with K neighbors. We use a quantitative measure of the magnitude of epistatic interactions (size of epistasis) and find that it is a monotonically increasing function of K. At high mutation rates more epistatic pairs are observed on the line of descent than expected, whereas at low mutation rates some epistatic interactions are selected against. Higher fitness is attained at an intermediate amount of ruggedness because both the height of the global peak and the mean selection coefficient per beneficial mutation increase with K. However, if K is too high, the longer waiting time between beneficial mutations counteracts those benefits. Increasing K transforms the fitness landscape from non-epistatic and smooth to epistatic and rugged, and increases the adaptive potential of the population. When the environment changes so that the population finds itself at a suboptimal location of an epistatic landscape, pleiotropy enables the population to increase fitness drastically with only a few mutational steps. Subsequently, crossing valleys between peaks in the fitness landscape requires deleterious mutations, which then become essential for adaptation by interacting with later mutations. Therefore, both deleterious and beneficial mutations contribute to adaptation, underscoring the benefit of initially deleterious mutations and epistasis in adaptive evolution. Increasing the number of interactions between loci on the one hand creates the opportunity for groups of loci to work synergistically as modules towards increased fitness. On the other hand, if the number of interactions is too high, the constraints they engender may be too severe for the adaptive process to take advantage of the potential.
 Sijmen Schoustra
Sijmen Schoustra
Properties of beneficial mutations
 Adaptation is one of the least understood processes in biology because it relies on beneficial mutations, which are often too rare to study. In my work I use experimental evolution in a filamentous fungus to address questions on the properties of these beneficial mutations. I will present a study with 120 experimental populations and a novel maximum likelihood method to infer the number and effect size of beneficial mutations substituted during adaptation, a process called an adaptive walk. This work shows that, in contrast to the gradualist view of adaptation dominant since the 1930’s, adaptive walks tend to be fast and short, with beneficial mutations of large effect substituted first followed by those of smaller effect. I will further present results of a study which addressed the effect of the level of dominance and of recombination of beneficial mutations by experimental evolution in otherwise isogenic vegetative haploid and diploid populations and shows that beneficial mutations are largely recessive. Recombination of multiple mutations yields genotypes that are more highly adapted than genotypes that fixed the same number of beneficial mutations in succession without recombination, which experimentally shows that recombination facilitates the crossing of adaptive valleys.
Adaptation is one of the least understood processes in biology because it relies on beneficial mutations, which are often too rare to study. In my work I use experimental evolution in a filamentous fungus to address questions on the properties of these beneficial mutations. I will present a study with 120 experimental populations and a novel maximum likelihood method to infer the number and effect size of beneficial mutations substituted during adaptation, a process called an adaptive walk. This work shows that, in contrast to the gradualist view of adaptation dominant since the 1930’s, adaptive walks tend to be fast and short, with beneficial mutations of large effect substituted first followed by those of smaller effect. I will further present results of a study which addressed the effect of the level of dominance and of recombination of beneficial mutations by experimental evolution in otherwise isogenic vegetative haploid and diploid populations and shows that beneficial mutations are largely recessive. Recombination of multiple mutations yields genotypes that are more highly adapted than genotypes that fixed the same number of beneficial mutations in succession without recombination, which experimentally shows that recombination facilitates the crossing of adaptive valleys.


