By Justin P. Hicks | jhicks3@mlive.com
Vaccines to protect against severe illness and death from COVID-19 started as the key to a return to normal, but they could wind up unlocking much more for the future of health care.
The mRNA vaccine technology used by Pfizer/BioNTech and Moderna for their respective coronavirus vaccines has been heavily touted by doctors and public health officials as a modern miracle of science and a means to revolutionize vaccine development.
“It’s hard to overestimate the impact this will have on human health,” said Nils Walter, a professor of biological chemistry at the University of Michigan who has studied mRNA for about 30 years. “It’s like introducing the iPhone when everyone had a flip phone.”
Beyond its uses for COVID-19, numerous mRNA vaccine candidates are being researched and undergoing clinical trials for the treatment of cancer, including pancreatic cancer, colorectal cancer and melanoma, as well as HIV, influenza, Ebola, Zika, and rabies.
“The same molecule from billions of years ago that gave us life is now coming back to what ails us today in life, which are viral infections and the terrible diseases like cancer,” Walter said. “It’s transformative.”
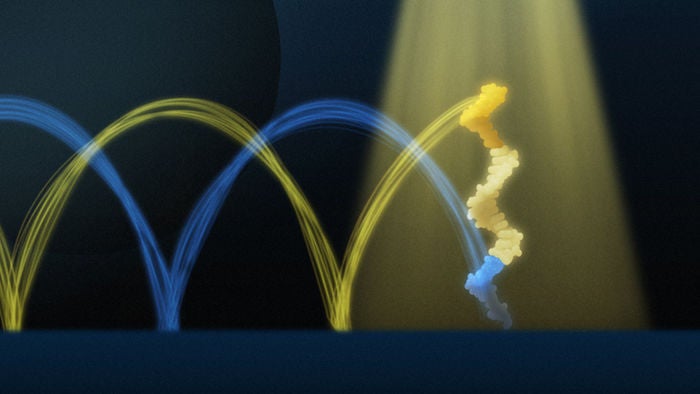
Vaccines using mRNA technology work by delivering instructions to the cells. In the case of COVID-19, those instructions are to create a protein that mimics the spike protein found in SARS-CoV-2, which triggers an immune response and the development of antibodies to defend against that virus.

Unlike previously used viral vector vaccines, mRNA vaccines don’t introduce live or dead virus into the body as a means of triggering antibody production.
The ingredients of the vaccine are broken down and discarded in a matter of days. The mRNA never enters the nucleus of the cell, and has no interaction with the cell’s DNA, health officials have repeatedly stated.
Dr. Liam Sullivan, an infectious disease specialist for Spectrum Health, expects mRNA technology to “revolutionize vaccine development.”
“This is just the beginning; we’re just scratching the surface here,” he said. “mRNA vaccine technology for treating diseases isn’t going anywhere. If anything, it’s going to get better and better and be used more widespread.”
Previous vaccines like the annual flu shot could take a year to develop, which made it challenging to predict which strains would be around by the next flu cycle. As a result, flu shot effectiveness has varied year to year.
With mRNA vaccines, that timeline can be cutdown 10-fold, Walter said.
“We can now operate at the same speed as the virus and therefore, viruses can no longer outmaneuver us and the vaccine can be made to specs in very, very short time,” he said. “It’s a totally different ball game.”
Continue reading